While Robert Koch asserted that infectious diseases were caused by bacteria, the existence of viruses —— biological entities even smaller than bacteria —— was not known at that time. “Virus” is a Latin word meaning “venom” and the perception of viruses as something bad has become firmly established thanks to the novel coronavirus that causes COVID-19. However, advances in genome analysis have led to the discovery that endogenous retroviruses and a variety of other viruses are involved in the evolution of many organisms. Viruses that do not fit conventional definitions have also been discovered. A new picture of viruses is emerging.
Special Feature 1 – The Unknown World of Viruses Shedding light on the role of viruses in biological evolution
composition by Rie Iizuka
illustration by Yoshihito Aoki
Koch, who in 1876 became the first person to successfully isolate Bacillus anthracis, the bacterium that causes anthrax, posited that infectious diseases were caused by bacteria. He was aided in his discovery by porcelain filters. Measuring just 0.2 µm, the fine pores in the porcelain were too small for bacteria to pass through, enabling Koch to capture them.
However, one cause of infectious diseases was found to pass through this apparatus. These agents were named “viruses,” from the Latin for “venom.” Viruses have been regarded as the enemy ever since that time.
Replicating only in the host’s cells, rather than self-replicating
Today, however, advances in molecular biology and techniques in other branches of science have shed light on the varying ways in which viruses behave, raising the question of what viruses actually are.
The textbook answer is that what all viruses have in common is their structure, consisting of nucleic acid in the form of DNA or RNA, which contains genetic information, surrounded by a protein shell called a capsid. However, they have neither the cell structure regarded as the basic unit of a living organism, nor the ribosomes required to synthesize proteins. Generally between 20 and 300 nm long (1 nm = 0.001 µm), viruses cannot self-replicate alone and are believed only to replicate in a host’s cells. While they share common constituent elements, in the form of nucleic acid and a capsid, the shapes of viruses vary. Some are spherical, while others are rod-shaped (Figure 1). Some even have an additional structure called an envelope. A viral envelope is a lipid bilayer membrane surrounding the nucleic acid and the capsid. However, rather than being produced by the virus, the membrane itself is formed by borrowing the host’s cell membranes and the like when the virus emerges from infected cells. Coronaviruses, too, have an envelope outside the capsid.
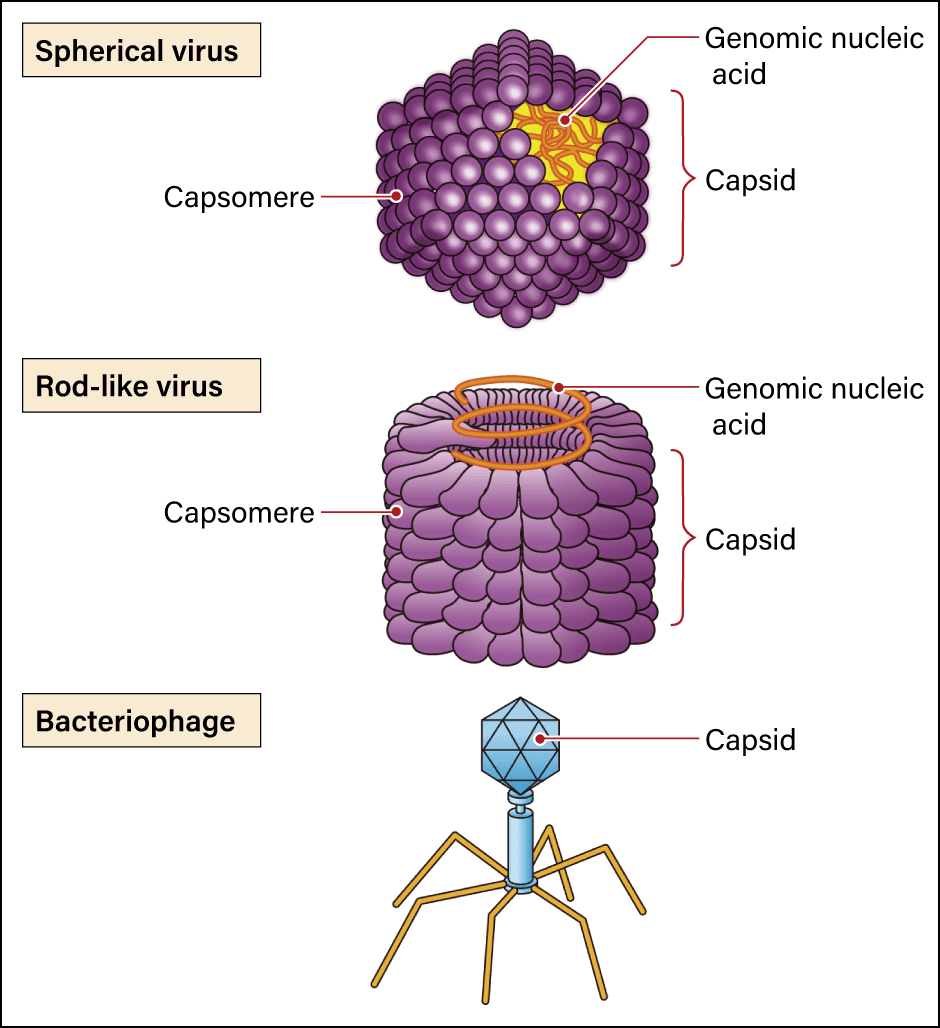
Figure 1. Viruses with various shapesThe simple word “virus” covers a variety of agents with different shapes, including spherical (icosahedral), rod-like, and filamentous viruses.
Earth is home to a variety of viruses, which are classified by the International Committee on Taxonomy of Viruses (ICTV), a body established in 1966. Among the agents recognized by the ICTV as belonging in the ranks of viruses are those consisting of RNA alone, without a capsid, and those that are about the same size as bacteria, both of which types have fallen outside the scope of textbook explanations until now.
Viruses without a capsid are often found in plants and fungi. They are considered to be members of the virus group because the features of the enzymes they use to replicate their own RNA are very similar to viruses that actually produce a capsid, so they are believed to be relatives. More specifically, capsidless viruses are thought to be viruses whose capsid atrophied or the ancestors thereof. For example, coronaviruses go through a cycle in which they emerge from cells in order to move into the cells of the next host. At such times, the capsid’s role is to protect the nucleic acid from the external environment, including ultraviolet rays and enzymes that degrade nucleic acid.
The criteria for defining viruses change with the times
However, it is thought most capsidless viruses basically do not leave the host’s cells once they have entered them, and therefore do not need a capsid. Each time a virus with new characteristics appears, the ICTV classifies it, so the criteria for defining viruses also change over time.
For example, the boundary between transposable elements (transposons) and viruses is quite vague. Transposon is the generic term for an element of a specific genome sequence in a nucleus that inserts itself into another position or makes more copies of itself. Transposons are found almost without exception in the genomes of organisms and those known as long terminal repeat (LTR) retrotransposons have a structure and genes very similar to retroviruses, one of the best-known of which is human immunodeficiency virus (HIV) (Figure 2).
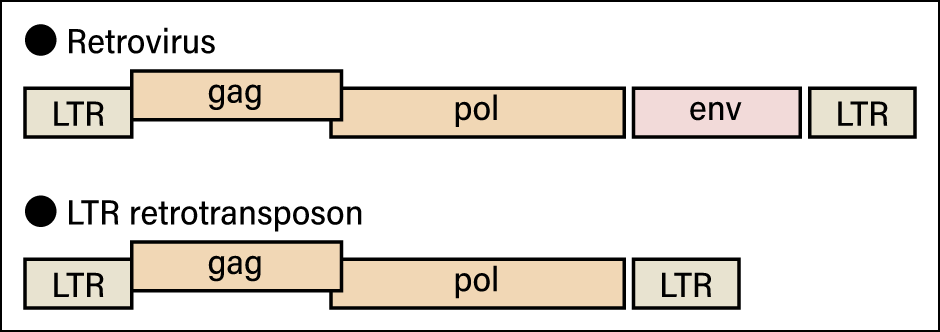
Figure 2. Schematic diagram of a retrovirus and an LTR retrotransposonThe only difference between the two is whether or not they have env, the gene involved in the creation of the envelope. All other parts are the same.
Although retroviruses have a gene that creates an envelope, typical LTR retrotransposons do not. However, as mentioned above, some viruses that do not leave cells have neither a capsid nor an envelope. At the same time, many retroviruses that have ceased to function because their envelope (env) gene has been destroyed are found in the genomes of living creatures. If we classified viruses and transposons according to whether or not they have an envelope, these retroviruses would end up being classified as transposons.
For example, when we compare coronaviruses, HIV, and LTR retrotransposons, HIV and coronaviruses are both classified as viruses because they cause infectious diseases, but if we look solely at their base sequences, any scientist would classify LTR retrotransposons and HIV as belonging to the same group, while coronaviruses would be quite far removed from them.
Moreover, DNA molecules called plasmids are found in bacteria and the like. As plasmids multiply in cytoplasm, independently from chromosomes, they have some similarities to viruses, but they also benefit bacteria, due to having genes resistant to antibiotics or assisting in gene transfer between cells, for instance. In a sense, they coexist with bacteria. Plasmids could be said to have adopted the strategy of sustaining themselves as cells proliferate.
Capsidless viruses were hitherto classified as viruses, due to their having a similar base sequence to agents defined as viruses. However, some plasmids also have gene sequences thought to share a common ancestry with virus genes. If we applied the same approach, these plasmids also would be classified as viruses.
Thus, it appears that what at first glance seem to be contradictions in the definition of viruses are related to the history of science.
The astonishing effects of viruses
Motivated by the role of viruses in causing disease in living creatures, research progressed, leading to the discovery of tobacco mosaic virus in 1898 and human yellow fever virus in 1901. The concept of plasmids as extrachromosomal genetic elements was propounded in the 1950s. Since then, attention has focused on their role as elements determining drug resistance and sex. The 1950s also saw scientists propose the existence of transposons as “jumping genes” controlling the movement of other genes. In other words, they were discovered as completely different things, but advances in technology and science gradually revealed what they have in common. Defining viruses is quite tricky.
While viruses are deemed not to be living organisms, some of them have astonishing ecological significance.
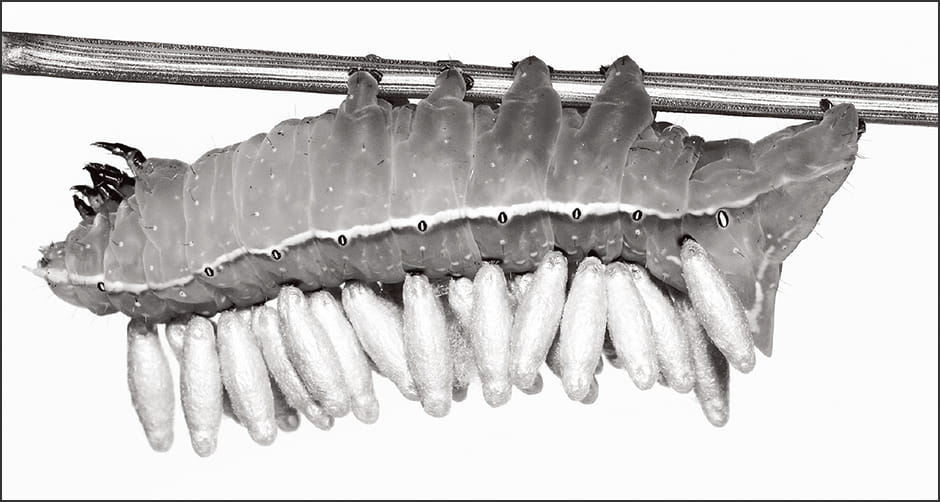
Braconids, a family of parasitic wasps, lay their eggs in a host called Mythimna (Pseudaletia) separata, which is a species of armyworm. When the braconid larvae emerge, the host armyworm begins to exhibit visibly peculiar behavior (Figure 3).
 (Photograph courtesy of Shinji Sugiura)
(Photograph courtesy of Shinji Sugiura)
Figure 3. Parasitic wasp larvae emerged from a hostThe parasitic wasps (Microgastrinae) on the back of the host (Amphipyra monolitha surnia) become pupae. The host exhibits behavior suggesting that it is controlled by a virus.
Whereas M. separata is a nocturnal species, it starts to linger around plant leaves during the daytime, as though preparing for the emergence of the braconid. The braconid exits its host by breaking through the surface of the armyworm’s body and forms a chrysalis on the leaf. Once the braconid has fully emerged, the M. separata summons up the last of its strength to move away from the leaf and dies elsewhere. This final action by the host is said to be carried out under the control of the braconid, to ensure that the armyworm’s decaying remains do not contaminate the chrysalis.
Although the mechanism behind this startling behavior on the part of the host is not yet understood, a family of viruses called polydnaviruses plays a major role in the parasitization of M. separata by braconids. Polydnaviruses are present in the ovaries of female parasitic wasps and are injected into the host when the wasp lays its eggs, serving to exercise powerful control over the host’s immune system. As insects have an innate immune system, a braconid trying to parasitize one ought to be eliminated by the host’s immunity. However, the polydnavirus prevents the host’s immunity from functioning by such means as promoting apoptosis (programmed cell death) in the host’s immune cells and impeding the activation of proteins that play an important role in the immune system. Furthermore, by controlling hormone secretion in the host insect, the virus serves to maintain it in a state beneficial to the parasitic wasp. What is even more interesting about polydnaviruses as viruses is that their viral particles do not have the genes required to produce a capsid or even to replicate.
So how do they maintain their physical form as viruses and replicate? In fact, research has shown that the proteins required for this purpose are provided by the parasitic wasp. The genes the polydnavirus needs to replicate are found in the parasitic wasp’s genome and trigger the provision of proteins such as the enzyme replicase. That is the indescribably strange tale of how a parasitic wasp keeps a virus in its genome and uses it to infect M. separata.
This is not the only example of its kind. The decoding of various genomes has resulted in the discovery that many organisms have virus-derived nucleic acid extensively inserted into chromosomal DNA within their nuclei. Endogenous retroviruses (ERVs) are a typical example.
In recent years, a number of discoveries have been made regarding important functions acquired by humans as a result of ERVs. A leading example is the syncytiotrophoblast, which is crucial to placenta formation in humans (Figure 4). A protein called syncytin derived from the ERV env gene is used in the formation of the syncytiotrophoblast. Allogeneic substances —— those not belonging to the individual —— are eliminated by the immune system in humans and other animals. However, the presence of the syncytiotrophoblast means that an individual can carry in its womb a fetus, which is a prime example of a foreign substance. This membrane serves to protect the fetus by allowing oxygen and nutrients to pass through in the blood supplied to the fetus from the placenta, while preventing the passage of lymphocytes and other immune cells that attack “foreign substances.”
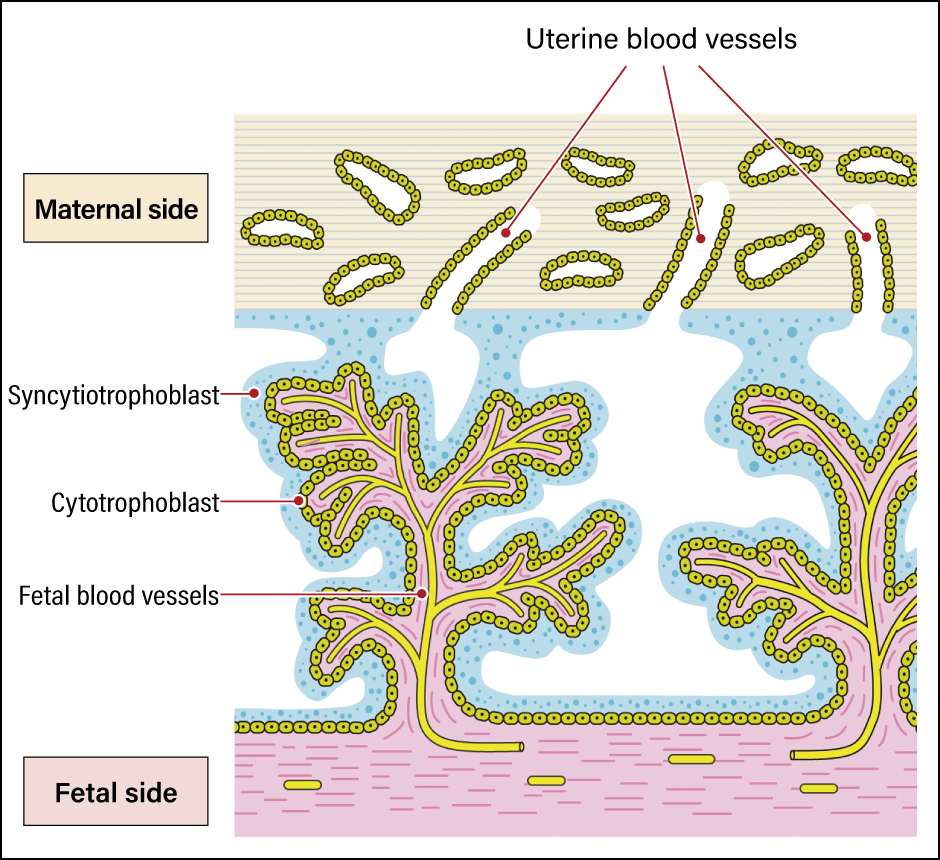
Figure 4. The syncytiotrophoblastThe power of viruses was leveraged to create a mechanism that protects the fetus by fusing cell membranes.
Usually, leukocytes and other immune cells can invade blood vessels by slipping between cells to attack their enemies. However, the syncytiotrophoblast has a structure more like cells fused together than a membrane and therefore has no gaps between cells, so immune cells cannot slip through it. In viruses, the syncytin that produces this structure functions by fusing the virus’s own envelope with the cell membrane of the infected organism, thereby promoting infection. In other words, the advanced mechanism of the human placenta was established when genes playing a role in cell membrane fusion, which viruses originally had for the purpose of infection, were incorporated into the human genome.
Viruses involved in advanced functions
One well-known human gene derived from viruses is skin aspartic protease (SASPase), which gives skin moisture retention properties. Organisms that emerged onto land in the process of evolution need to maintain moisture in their skin. Proteins need to be broken down in the course of creating the skin’s moisture-retaining layer and ERV-derived SASPase is used as an enzyme in this process. Interestingly, one example suggesting that virus-derived genes are involved in biological evolution is the fact that SASPase is only found in mammals, which have skin without scales.
Discovered in 2006 or thereabouts, activity-regulated cytoskeleton-associated protein (ARC) is a protein that plays a part in aiding the transmission of RNA between neurons. Fundamentally speaking, ARC is a protein involved in the formation of virus particles in which RNA is packaged, and it is thought to assist in the transmission of RNA between cells by leveraging its original function. Thus, viruses are even involved in advanced functions of this kind.
Human genome analysis has revealed that LTR retrotransposons and other transposable elements account for as much as 45% of the entire genome. While viral fragments had been identified scattered here and there in the human genome, they were initially deemed to be “junk DNA” without any function. However, recent research has shown that these viral fragments play an important role as gene regulatory sequences.
For example, the renowned Yamanaka factors are four transcription factors introduced to create induced pluripotent stem (iPS) cells. In doing so, rather than making four genes function, each of these four transcription factors control a number of genes. One of the Yamanaka factors, the transcription factor called Oct-4, is bound to a sequence derived from ERV1, a type of ERV in the human genome. As ERV1 is repeated many times in the genome, if Oct-4 needed to regulate 100 genes, for example, it could simultaneously control 100 different types of gene in different places, so long as ERV1 was located near those genes.
In the human body, too, when there is a need to activate a large number of genes, using an ERV scattered throughout the genome as a binding sequence could potentially enable the factor bound to them to function like a main power switch. In other words, switching it on and off would effectively light up all 100 genes —— that is to say, express those genes. This would make it easy to form a mechanism for controlling downstream genes en masse, like switching off the lights at certain times. Some researchers are starting to take the view that ERV-related repetitive sequences in the genome could be significant in providing this kind of mechanism.
Having started from the discovery that the substances syncytin, SASPase, and ARC play an important role in creating what we know as humans, we are now beginning to see that human genes are structured and networked by the virus sequences scattered throughout the genome. While viruses pose a threat of infectious disease, they are also an agent essential to our evolution.