Covering about 70% of the earth’s surface, the ocean is home to a diverse array of organisms that significantly affect our planet’s ecosystem. These include microscopic phytoplankton, which play a major role as a carbon sink. This means marine phytoplankton are a crucial factor influencing climate. And it is virtually certain that giant viruses are one of the factors regulating phytoplankton. Unraveling the relationship between these giant viruses and phytoplankton is likely to make a substantial contribution to future measures to combat climate change.
Special Feature 1 – The Unknown World of Viruses The giant viruses regulating marine phytoplankton
composition by Takakazu Kawasaki
The ocean covers about 70% of the earth’s surface. Containing vast ecosystems, they are a source of aquatic resources and serve as a sink absorbing anthropogenic carbon dioxide (CO2) emissions. Just 1 ml of surface seawater from a typical open ocean contains anywhere from at least several thousand to tens of thousands of eukaryote cells, hundreds of thousands to several million prokaryote cells, and tens of millions to several hundred million virus particles.
- *1 Eukaryotes: Organisms whose cell nucleus houses DNA. They include unicellular and multicellular animals, plants other than cyanobacteria, and eukaryotic fungi.
- *2 Prokaryotes: Organisms consisting of cells without a nucleus (prokaryotic cells). bacteria and archaea fall into this category.
The viruses determining marine material flows
The most important primary producers (autotroph) in the ocean are microscopic phytoplankton. Recent studies involving the genetic analysis of marine microorganisms have revealed that phytoplankton fix atmospheric CO2 via their interaction with marine giant viruses, transporting it to the ocean depths and even affecting the weather (Figure 1).
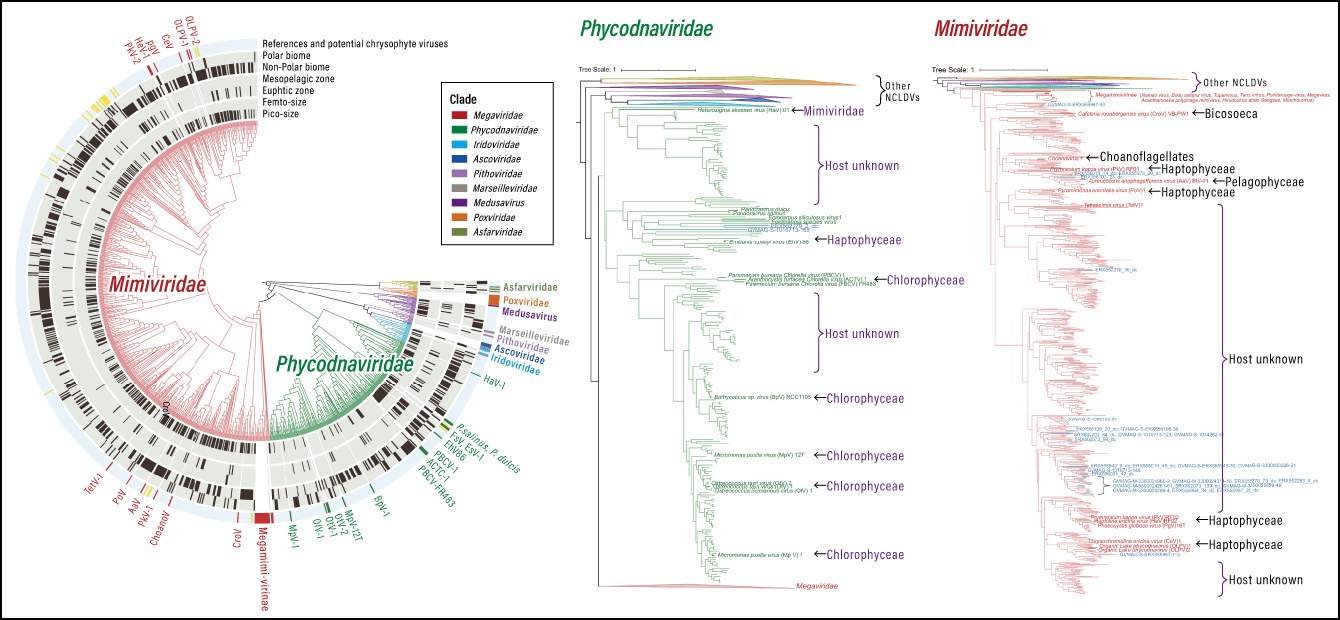 (Phylogenetic tree for giant virus PolB sequences: Constructed from the reference sequences for known phylotypes and 905 long (≥700 amino acid) environmental sequences)
(Phylogenetic tree for giant virus PolB sequences: Constructed from the reference sequences for known phylotypes and 905 long (≥700 amino acid) environmental sequences)
Figure 1. Phylogenetic affiliations of nucleocytoplasmic large DNA viruses (NCLDVs) detected in marine metagenomesThe phylogenetic tree of giant viruses identified from marine metagenomes. Particularly high levels of diversity were found in the tree for Phycodnaviridae and Mimiviridae (especially Mesomimivirinae). Many of the branches are distantly related to known hosts, suggesting the existence of a more extensive host clade.
While the debate about whether or not viruses are living organisms continues, the fact is that viruses can neither produce or metabolize energy alone, nor propagate themselves. To proliferate, viruses need to infect a living host and use the molecular machinery via which cellular metabolism and protein synthesis in the host takes place. In most cases, viruses that have proliferated in a host are released into the environment after destroying the host’s cells. In other words, for unicellular microorganisms, viral infection means death. In the ocean, organisms are affected by viral infections at a wide range of trophic levels, from bacteria to microscopic phytoplankton and zooplankton, all the way up to fish. Some viruses can even influence the community dynamics of microorganisms by modifying the host’s physiological functions.
If the host cell is destroyed by a viral infection, the organic carbon, nitrogen, phosphorus, and other trophic resources contained in the cell are released into the environment, where they may be incorporated into other organisms via decomposition by heterotrophic bacteria or condense into particles that sink to the ocean depths.
- *3 Heterotrophic bacteria: Bacteria requiring a source of organic carbon to grow.
The ocean’s layers are broadly classified by depth into the surface layer, or epipelagic zone (0–200 m), the mesopelagic zone (200–1,000 m), and everything below that. In the euphotic zone —— the part of the epipelagic zone between 50 and 150 m deep that is reached by sunlight —— phytoplankton use CO2 from the atmosphere dissolved in seawater to carry out photosynthesis. Phytoplankton generate organic matter such as glucose (C6H12O6) from CO2 by means of photosynthesis. Some phytoplankton produce calcium carbonate (CaCO3) from carbonate ions (CO32-) dissociated from CO2 and calcium ions (Ca2+) in seawater. As CO2 is generated by an equilibrium reaction in the carbonate system when these reactions take place, calcification could also be described as a reaction that emits CO2, in the opposite manner to photosynthesis. In other words, if we investigate the phytoplankton species and quantity of organisms present in the marine surface layer, we can deduce the properties of the carbon cycle in that body of water.
Analyzing marine genetic data
We conduct research into giant viruses living in the oceans that have comparatively large viral particles and genomes. “Giant virus” is not the official name for a viral taxon. There is no clear boundary for defining such viruses by genome size or particle size, and their visible characteristics also vary from one species to another. Generally speaking, most of the viruses called giant viruses have a particle size of at least 200 nm and a genome size of at least 100,000 base pairs, but the category does also include smaller viruses.
All giant viruses are deemed to belong to the phylum Nucleocytoviricota of viruses. However, as a succession of new species of giant viruses have been isolated from all kinds of environments in recent years, many giant viruses are deemed to be giant viruses in substantive terms, without waiting for their official scientific name to be conferred by the International Committee on Taxonomy of Viruses (ICTV).
The first giant virus to be discovered was Acanthamoeba polyphaga mimivirus, which was the subject of an article published in 2003 by a team led by Bernard La Scola, a microbiologist at Aix-Marseille University in France. In fact, Acanthamoeba polyphaga mimivirus had actually been discovered in 1992, but as it was large enough to be observed with an optical microscope, it was thought to be a bacterium.
When next-generation sequencers were subsequently developed, genome sequencing progressed rapidly. The genomes of organisms are usually at least several million bases long, so decoding them required the ability to read a large volume of sequences. The next-generation sequencers in operation today can read several billion sequences at once, so we can now cheaply obtain sufficient information to determine an organism’s genome sequence. However, shotgun sequencing, which is the main technique used at present, reads short DNA sequences with the aid of restriction enzymes or ultrasound. Consequently, the sequences have to be joined up on a computer to restore them. It is similar to the task of joining up scraps of paper from a shredder to reconstruct the original document.
Using a next-generation sequencer has made it possible to decode the genes of microorganisms quickly and comprehensively, without the need to classify them into one of what are said to be tens of thousands of marine plankton species or isolate microscopic plankton. And this has made it possible to guarantee the objectivity and quantitative properties of data on microorganism and viral communities, to some extent. While there are still some issues to be addressed, progress is being made in creating a database of observation data.
Recent years have seen the development of even more advanced long-read sequencers, enabling long DNA sequences to be read without the need to break them into fragments. These sequencers are likely to become the main apparatus used in genetic research in due course, and I myself use them in my research into marine ecosystems.
The key functions of marine ecosystems
As part of an international marine exploration project called Tara Oceans, a research expedition covering nearly all the seas on the planet was conducted between 2009 and 2013 to study giant viruses. The main focus of the survey was genetic data such as DNA and RNA in seawater. A total of 120 locations were surveyed, from the Antarctic to the Arctic Ocean.
I was a graduate student at Hokkaido University at the time and was invited to join the voyage by the project coordinator, who I had happened to meet at an international academic conference, but I was unable to take part, due in part to my other research. However, after transferring six years later to Kyoto University’s Institute for Chemical Research, where I now work, I took on responsibility for analyzing the data from the project, which makes me think it was somehow meant to be.

Assistant Professor Endo (left) collecting samples in the Antarctic Ocean.
The DNA derived from approximately 300 microbial communities gathered by the Tara Oceans expedition and semi-comprehensively read by the next-generation sequencer is metagenomic data. Currently, the 370 pieces of marine metagenomic data gathered by the Tara Oceans expedition was used to build a marine genetic catalog containing approximately 47 million gene sequences. We searched this gene catalog for the sequences of family B DNA polymerase (polB) gene, which is a marker genes for giant viruses, and detected approximately 7,000 varieties (phylotypes) of giant viruses, including many unknown phylotypes (Figure 1).
- *4 Family B DNA polymerase: An enzyme that uses single-stranded DNA as a template to synthesize double-stranded DNA.
In the oceans, seawater is mixed into the epipelagic zone down to a depth of around 50 to 100 m, but there is hardly any mixing at all below that depth. For substances to move from the epipelagic to the mesopelagic zone, they must either settle under their own weight due to gravity or be carried there due to vertical migration of fish or other large organisms.
However, looking at the data from our analysis, there are many observation sites where similar viruses are found both in the epipelagic and mesopelagic zones. It seems as though viruses travel down from above, but even comparatively large giant viruses are not heavy enough to fall under their own weight. For giant viruses to move downward, they must infect a bigger, heavier host. For example, it is reasonable to think that aggregates of plankton and the like are involved in the biological carbon pump, which carries carbon compounds to the lower layers of the ocean.
Plankton that have been stressed or killed by a viral infection tend to form aggregates. When phytoplankton that have grown due to photosynthesis in the epipelagic zone sink after being killed and forming aggregates, the resultant CO2 that dissolves into seawater from the atmosphere is sequestered in the ocean depths over a period of several hundred to several thousand years. This function is called the biological carbon pump. It is estimated that, without this mechanism, the concentration of CO2 in the atmosphere would be double the current level. Thus, the marine ecosystem fulfills an important function in maintaining the climate in a stable state.
Factors regulating phytoplankton
Researchers in Canada have propounded the highly interesting theory that, by killing hosts and promoting their aggregation, viruses are involved in the efficiency of the biological carbon pump. In addition, when the team at our laboratory analyzed the Tara Oceans data, there appeared to be a very strong correlation between the structure of virus communities and the efficiency of the biological carbon pump.
We also obtained results suggesting that viruses that infect major marine phytoplankton, namely diatoms and haptophytes, held the key to the efficiency of the biological carbon pump (Figure 2).
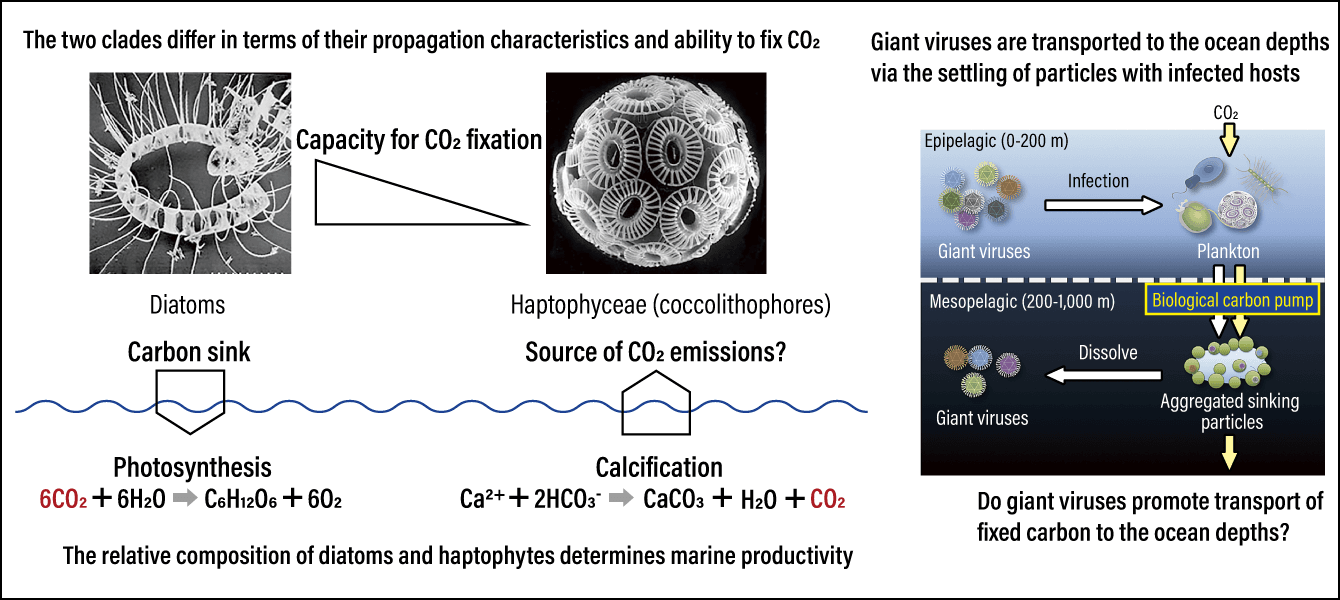
Figure 2. Diatoms and haptophytes: major marine phytoplankton<Left> The major phytoplankton groups, diatoms and haptophytes, demonstrate substantial differences in terms of their ability to fix CO2. <Right> A study by Endo and his team revealed that the viruses that infect diatoms, haptophytes, and other major phytoplankton hold the key to the efficiency of the biological carbon pump, which exports CO2 in the ocean’s mesopelagic zone.
Diatoms and haptophytes affect the ability to fix CO2 in the ocean in contrasting ways. Diatoms engage in intense photosynthesis, lowering the CO2 partial pressure in the seawater and encouraging CO2 to dissolve into it from the atmosphere. At the same time, the clade coccolithophore that calcifies haptophytes reduces carbonate buffering in seawater in the calcification process, thereby creating an environment in which it is more difficult for CO2 to dissolve into the seawater from the atmosphere. In other words, oceanic regions where haptophytes predominate are environments with a poor capacity to absorb CO2 via photosynthesis. Even when investigating plankton fossils (microfossils) by era, we can see that diatoms were predominant during the cold glacial period, while many haptophyte fossils have been found dating back to the warm interglacial period.
Phytoplankton might possibly produce specific chemical substances to protect against infection by giant viruses. For example, dimethylsulfide (DMS) is produced when bacteria break down the organosulfur compound dimethylsulfoniopropionate (DMSP), which is a metabolite of algae. It has been reported that DMS has the effect of protecting hosts from viral infection.
Interestingly, DMS readily absorbs water, so if it is generated in large quantities, it is released into the atmosphere, where it forms condensation nuclei that become clouds. It has been proposed that, as clouds block sunlight, this could have the effect of cooling the earth. Haptophytes in particular are known to produce DMSP in abundance.
In highly productive marine areas with a lot of host plankton in the epipelagic zone, there are also many viruses capable of killing them. However, if the effectiveness of those viruses weakens and the ecosystem cycle stagnates, a process that could be described as plankton aging occurs, changing the composition of the organic matter released. There have been reports that this might inhibit cloud formation and rainfall.
As the earth’s climate system is very complex, it might be difficult at this point to draw a simple link between the function of marine viruses and global warming. However, the impacts of phytoplankton biomass and community composition on biochemical cycles are comparatively well understood. It is almost certain that giant viruses are a factor regulating phytoplankton communities. Accordingly, if light can be shed on the scale of their impact in quantitative terms across an extensive area, we believe it will be possible to discuss the extent to which viruses affect the scale and nature of future warming.
Projections suggest that if warming progresses, it will be increasingly difficult for the ocean’s epipelagic zone to mix with the zones below it, causing a gradual decline in the concentration of nutrients supplied from the ocean depths and a concomitant reduction in phytoplankton biomass. Consequently, unraveling the relationship between viruses and the biological carbon pump is also essential from the perspective of maintaining the ocean’s function as a CO2 sink in the future.
Accordingly, we are trying to verify the role of viruses in driving the biological carbon pump in the Oyashio regions in the Pacific, which is the marine area that currently makes the world’s biggest contribution to the biological carbon pump.