Honeybee populations are in decline. The fall in the number of European honeybees, which are the main species used in beekeeping, is particularly pronounced. The main causes with direct impact, according to scientist’s theories, are disease borne by varroa mites, and hornets, which are natural predators of honeybees. In contrast, the Japanese honeybee —— a subspecies native to Japan —— is resistant to varroa mites and employs a technique called the “hot defensive bee ball” to fight off hornets. Scientists say that genetic analysis of the Japanese honeybee could potentially be applied to its conservation and even to selective breeding of honeybees.
Special Feature 1 – The World of Honeybees Genome analysis of the Japanese honeybee could help prevent honeybee decline!
composition by Takeaki Kikuchi
Honeybee populations are declining worldwide. The fall in the number of European honeybees (Apis mellifera), which are well-suited to beekeeping, is especially marked. Among the contributory factors cited are pathogen epidemics, climate change, a lack of flower resources, invasion by natural predators, domestication bias, and pesticides. These factors are believed to be intertwined in complex and varying ways. The extent of decline among Japanese honeybees (Apis cerana japonica) is unclear. However, unlike European honeybees, Japanese honeybees are characterized by a resistance to pathogens, among others, so they are attracting growing attention.
Comparison of the whole genome sequences in the Japanese honeybee
A subspecies of the Asian honeybee (A. Cerana), the Japanese honeybee is Japan’s only wild species of honeybee, which inhabits areas in and southward of the island of Honshu. Japanese honeybees are distributed across Honshu, Shikoku, Kyushu, and some of Japan’s outlying islands, and previous studies regarded all populations in the Japanese archipelago as being genetically homogeneous. However, as the Japanese archipelago stretches a long distance from north to south, there are variations in many of the conditions that affect honeybee survival, including temperature, altitude, precipitation, and snowfall. Accordingly, I hypothesized that environmental conditions might promote genetic differentiation and unique forms of evolution in Japanese honeybees. Given the extensive distribution range between the northern limit in Aomori Prefecture and the southern limit in Amami-Oshima Island, I thought there might be adaptive populations that differed from each other.
Taking a different approach from previous studies, I attempted to employ the latest whole genome sequencing. As a result, I discovered that, broadly speaking, Japanese honeybees fall into three genetically distinct geographic regions.
Previous studies based their comparisons of regional variation on morphological and genetic (rather than genomic) approaches.
The morphological approach generally analyzes wing vein length, based on a classification method called cluster analysis, which involves collecting similar objects from a mixture, and grouping them into clusters based on similarity. However, no regional characteristics have been identified using this approach. Moreover, in a previous study that compared the body size of Japanese honeybees from Iwate Prefecture, Kagoshima Prefecture, and Amami-Oshima Island, a significant difference was identified between bees from mainland Japan and those from Amami-Oshima Island, but no significant difference was observed between bees from Iwate Prefecture and those from Kagoshima Prefecture, despite the substantial distance between the two regions.
Studies using the genetic approach, meanwhile, have focused on comparisons of the mitochondrial DNA D-loop region. As base substitutions are frequently observed in the non-coding region called the D-loop, which is found in DNA located in cell organelles called mitochondria, the pace of evolution is said to be faster in this region than in others. However, no clear regional variations emerged even from these comparisons.
Previous studies compared the polymorphisms in just a few sites on the genome, so only limited traits and mutations could be compared. Updating these comparisons to a whole genome scale makes it possible to spot polymorphisms in millions of sites and enables us to assign meaning to mutations by investigating all mutations, rather than turning our attention to only specific traits or variants.
In fact, a number of whole-genome-scale studies of the regional variation of insects in the Japanese archipelago have already been carried out. For example, one study found that the Genji firefly (Luciola cruciate) is divided into a number of regional groups, ranging from Kyushu in the south of Japan to Tohoku in the northeast, and that there is a difference in their flash patterns between east and west Japan.
Difference in genetic characteristics among the three geographic regions
Our research team sequenced the whole genomes of 105 individual Japanese honeybees from across the three main Japanese islands, ranging from Aomori Prefecture to Kagoshima Prefecture (Figure 1). From these whole genomes, we identified single-nucleotide polymorphisms (SNPs) —— indicating differences in the DNA sequence —— at around 1.27 million locations, which we then compared in detail. Using this information, we estimated their genetic structure.

Figure 1. Distribution of the Japanese honeybees subject to genome analysisWith the cooperation of the National Agriculture and Food Research Organization (NARO) and beekeepers, the research team analyzed the genomes of 105 Japanese honeybees from across Japan, from Aomori Prefecture to Kagoshima Prefecture. The majority of the colonies raised by apiarists were derived from wild colonies in each region.
First, we used ADMIXTURE analysis to divide honeybees into genetic groups with similar genetic compositions, and then visualized it. This analysis showed obvious differences between Japanese honeybees and Asian honeybees from China. Japan archipelago is said to have split from the continent around 100,000 years ago. We can probably say that the genetic structure of the Japanese honeybee had been formed over the millennia since then.
From our analysis, it emerged that Japanese honeybee populations on the Japanese archipelago can broadly be categorized into three geographic regions: Northern (Tohoku, Kanto, and Chubu districts), Central (Chugoku district), and Southern (Kyushu district). We can see that, whereas the Northern region is dominated by blue, the Kinki and Shikoku districts are a mixture of multiple colors, while green is predominant in the Central region and orange in the Southern region (Figure 2). It was thanks to large-scale genomic information that the existence of this diversity became apparent.

Figure 2. Distinct genetic structure of Japanese honeybees confirmed in the Japanese archipelagoAs shown in the upper left-hand diagram, there were obvious differences between Asian honeybees from China and Japanese honeybees. Clear differences observed between the bees in three regions: the Northern region, encompassing the Tohoku, Kanto, and Chubu districts; the Central region, consisting of the Chugoku district; and the Southern region, consisting of the Kyushu district. The Kinki and Shikoku districts had a mixture of the three.
“Possible non-native” individuals were found among the bees sampled. For example, one honeybee from Fukushima Prefecture in the Northern region had the same genetic composition as Southern honeybees. When we investigated that bee’s “ancestry,” we discovered that it had actually been introduced to Fukushima Prefecture from Kumamoto Prefecture. The majority of the other exceptions were also suspected of having been imported from other regions by humans.
This suggests that, when individuals are transported between different regions, imported bee colonies might not be able to adapt to their new environment. Many people keep Japanese honeybees as a hobby and such hobby apiarists interact with each other, sometimes exchanging or selling bees. In our survey, we were able to detect situations where bees had been sent to a region with different circumstances from those to which they had originally adapted.
We tried to identify the genetic regions under natural selection in each of the three genetically distinct populations. Moreover, we investigated such selective regions affected by each environmental factor.
Looking first at differentiation on a population basis, we sought to detect geography-specific genetic regions, such as genetic regions that are similar in Northern and Central populations but differentiated in the Southern population, and genetic regions that are similar in Southern and Central populations but differentiated in the Northern population. This analysis would enable to find out which genetic regions contributed to adaptation in each of the three genetically distinct geographic regions.
As a result, we identified Shaker (potassium voltage-gated channel protein Shaker isoform X3) as a genetic region specifically differentiated in Southern populations. It is a gene potentially associated with abnormal behavior, lifespan, and sleep in fruit flies (Drosophila) (Figure 3). Nine genes, including GDPGP1 and CBS, were found in the specifically differentiated genetic region in Central populations. GDPGP1 controls glycogen quantity, while CBS is important in methionine metabolism and is associated with energy metabolism. Six genes, including NPC2 and HYPK, were specifically differentiated in Northern populations. The expression level of NPC2 is reported to differ in the event of infection with a pathogen, while HYPK is associated with apoptosis (cell death), and therefore could potentially be linked to immune function.
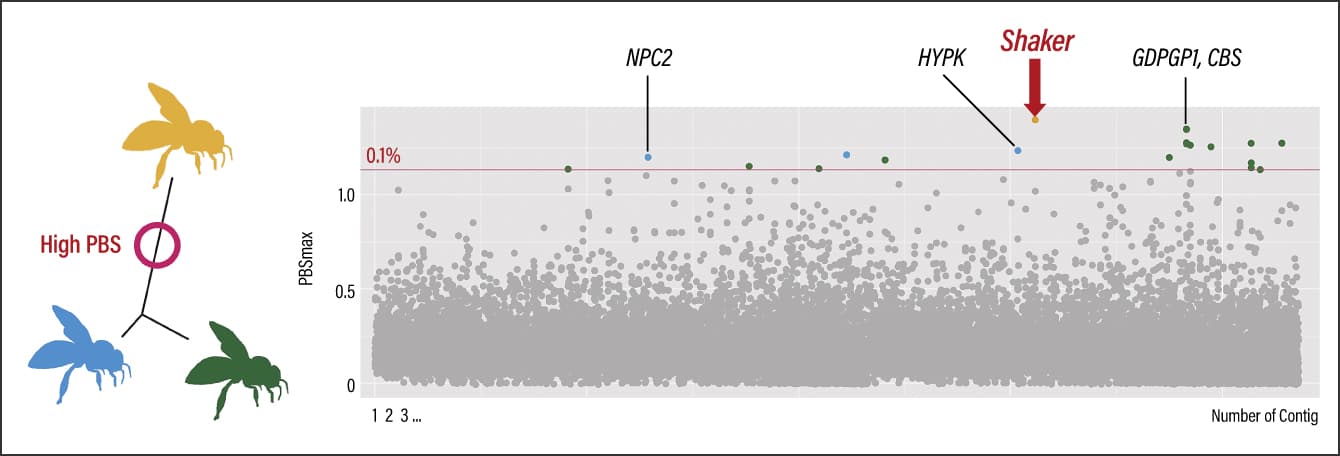
Figure 3. Genetic region of specific differentiation in Southern populations (Shaker)Population branch statistic (PBS) is a statistical measure used as an indicator of the degree of genetic differentiation. The whole genome was divided into specific ranges and the scores were calculated for each. A total of 25 genes were detected from the outlier window region in the upper 0.1% range. Specific differentiation was found in the region surrounding Shaker in the Southern region populations.
We also investigated the correlation between genes and environmental differences, specifically temperature, snowfall, precipitation, and hours of sunlight (Figure 4). This involves, for example, detecting patterns in which the genotype changes like a gradation as the average temperature at the location where the sample was collected rises. Although our analysis on the basis of correlation with these environmental variables succeeded in detecting around 70 candidate genes, none had any points in common with the 25 genes associated with differentiation on a population basis. This suggests that Japanese honeybees adapt to the specific environment in each region, making it unfeasible for them to simply move further north if temperatures rise in the future.
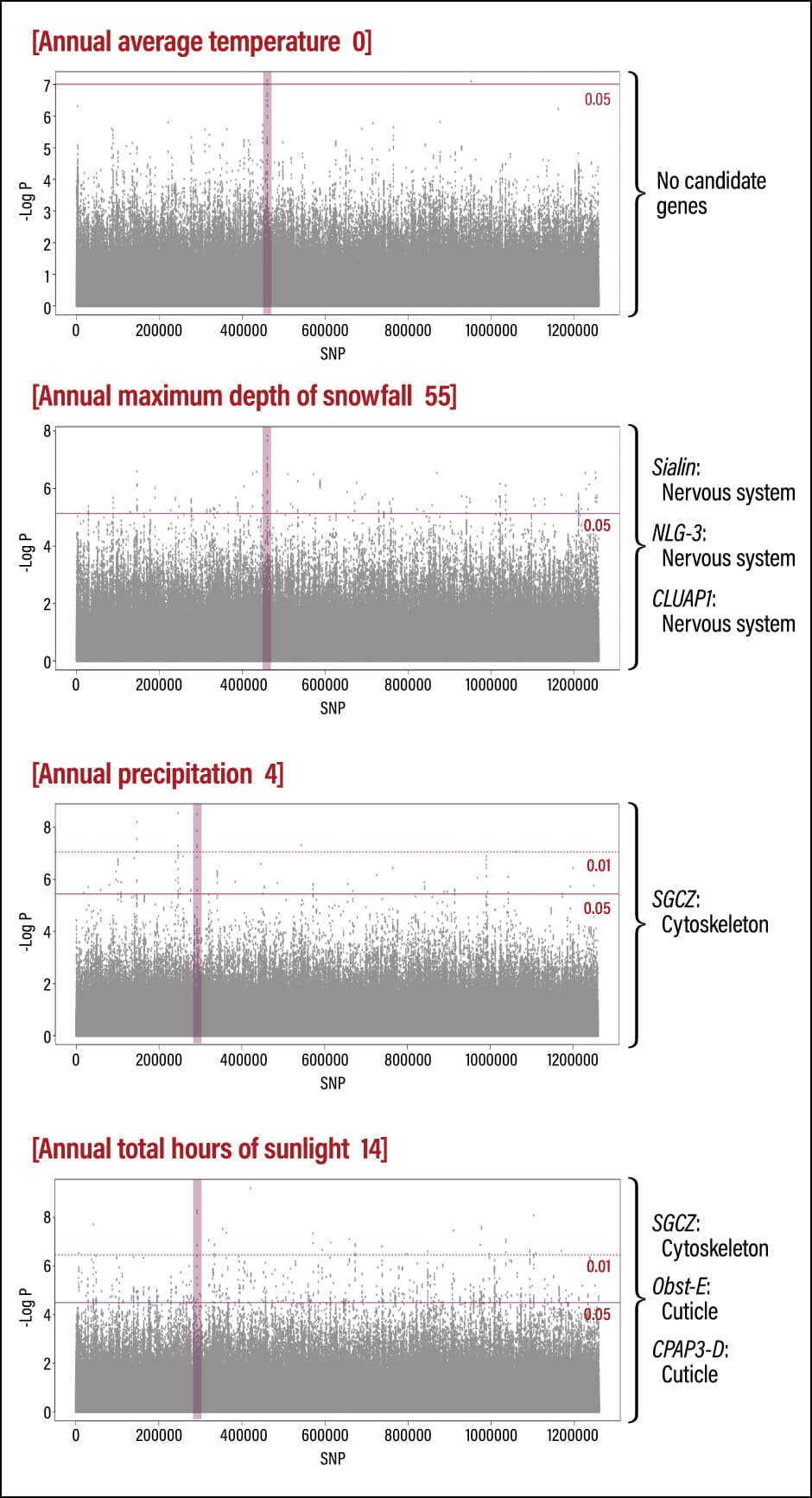
Figure 4. Correlation between environmental differences and genesWe sought to detect candidate genes correlated to environmental variables. However, we could not identify a candidate gene correlated to annual average temperature. On the other hand, there was a correlation between snowfall depth and several candidate genes associated with the nervous system, namely Sialin, NLG-3, and CLUAP1.
Prospects for honeybee evolutionary genomics research
The above is what we have learned about the genetic differentiation and local adaptation of the Japanese honeybee. If a simple system for detecting which of the three regional population groups a bee comes from can be developed, it will hopefully become possible to impose appropriate restrictions on honeybee movement and diagnose regional suitability. Investigating to which specific environments candidate genes associated with local adaptation are correlated and the specific ways in which they are involved in adaptation (for example, activity level, flower preferences, and resistance to disease in a specific environment) could enable us to contribute to both the conservation and the utilization of Japanese honeybees.
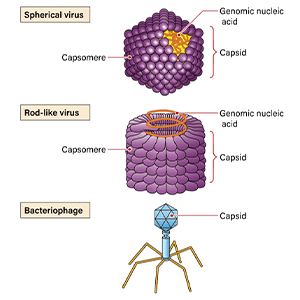
The leading example of a disease found in European honeybees, which are the main species used in beekeeping, is varroosis, which is caused by the varroa mite (Varroa destructor). This parasitic mite lives off bee larvae and pupae, causing a developmental disorder by sucking their bodily fluids. Varroa mites are also hosts for at least five viruses, including deformed wing virus. When larvae infected with deformed wing virus emerge, their wings do not grow normally and they die. The resistance of Asian honeybees, including Japanese honeybees, to the varroa mite is cited as one reason why they are resistant to disease.
The varroa mite was originally a parasite of the Asian honeybee. It then transferred to European honeybees introduced to Asian countries for the purpose of beekeeping. Japanese honeybees and European honeybees compete with each other when foraging in places with flowers; Japanese honeybees have even been observed entering European honeybee hives and stealing nectar. The interaction of the two honeybee species in this way enabled varroa mites to move from one to the other. It is thought that the varroa mites accompanied such honeybees when they were subsequently imported or exported internationally, spreading across the globe as a result. The way in which they moved has also been revealed by tracing their sequences. A research team led by Professor Stephen J. Martin of the University of Salford in the UK has published articles on the genetic analysis of honeybee pathogens.
The existence of natural predators has been cited as a contributory factor in the decline of European honeybees. One such predator is hornets. Japanese honeybees use a defensive measure called a hot defensive bee ball to resist hornets that approach their hives. Several hundred Japanese honeybees will surround a single hornet, as though engulfing it; they then move their flight muscles simultaneously to generate heat, raising the temperature inside the bee ball to 45-47°C. After 20-40 minutes inside the ball, the hornet dies from the heat. The difference between the fatal temperature for hornets and that for Japanese honeybees is about 3°C; Japanese honeybees leverage this temperature difference to fight off hornets.
Although European honeybees exhibit a similar behavior, they are attacked and destroyed before they can form a proper hot defensive bee ball. Hornets principally live in Asia. The difference between Asian honeybees, which have always lived in Asia, and European honeybees, which traditionally inhabited regions with few hornets, is apparent in this attribute.
The detailed sequencing of the European honeybee genome is already progressing, ahead of work on the Japanese honeybee genome. If, in due course, we can use genome analysis to compare the characteristics of the Japanese honeybee with those of the European honeybee and, for example, identify genes providing powerful resistance against mites, it might become possible to use selective breeding to develop European honeybees that can withstand mite-related diseases. In addition, identifying genes crucial to the formation of the hot defensive bee ball might open up the possibility of instilling in European honeybees attributes that could help them stand up to hornets.
Humans receive significant benefits from honeybees. These go beyond simply eating the honey and royal jelly that they produce. A major benefit is pollination. Honeybees are said to support one-third of global food production. Amid fears that future global population growth could lead to food shortages, honeybee decline will be a problem of acute concern. For this reason, too, I feel it is vital to use genome analysis and other advanced technologies to learn about the Japanese honeybee in greater detail.